NextGen Omics 2024

23 - 25 October 2024 | London, UK
NextGen Omics 2024 features 4 outstanding programmes, bringing together Europe’s most successful omics research experts under one roof.

Explore NextGen Omics 2024: Unlocking Future Insights in Multi-Omics Research
Join 700+ leaders, experts and researchers at our flagship NextGen Omics 2024, featuring the 16th Annual NGS & Clinical Diagnostics Congress, 12th Annual Single Cell & Spatial Analysis Congress and 6th Annual Digital PCR & Liquid Biopsies Congress and CRISPR-Based Medicine Congress
16th Annual Next Generation Sequencing & Clinical Diagnostics Congress
An intensive meeting with presentations from industry-leading researchers on the latest innovations in clinical genomics, multi omics, clinical diagnostics & omics bioinformatics. This must attend event will discuss the current state of the industry, looking at innovations in diagnostics development, clinical genomics, multi omics data analysis, integration & harmonisation. From new and exciting technologies to the latest in products & services, our conference will bring together leading research institutions & companies for engaging discussions, knowledge sharing and focused networking.
- Hear from and meet with the key innovators in multi-omics. 2023 attendees included: Senior Director, Bristol Myers Squibb; Head of Molecular Biology, Cerevance; and Vice President, Evaxion Biotech
- Take a deep dive into diagnostics approaches for infectious & non-infectious diseases, multi-omics in precision medicine, and microbiome profiling & analysis. Presentations will cover CRISPR-based diagnostics approaches, optimising therapeutic design for cell & gene therapies, and host-microbiome interactions
- Discuss the pioneering developments in clinical genomics and multi-omics in cancer discovery. Hear about newborn screening, ML-driven target identification, and tools & approaches to understand tumour heterogeneity
- Over 100 case studies, presentations, and panel discussions focused on key issues such as clinical genomics, multi-omic data integration, NGS applications in the clinic, diagnostics development, and advanced bioinformatics
Agenda at a Glance
Day One | Track 1: Diagnostics Approaches for Non-Infectious Diseases in Pharma
Day One | Track 2: Diagnostics Approaches for Infectious Diseases in Pharma
Day One | Track 3: Bioinformatics Approaches, Methodologies & Technologies
Day One | Track 4: Leveraging Multi-Omics in Precision Medicine in Pharma
Day One | Track 5: Optimising Cell & Gene Therapies with Multi-Omics Approaches in Pharma
Day One | Track 6: Microbiome Profiling & Analysis
Day Two | Track 1: Clinical Genomics
Day Two | Track 2: Multi Omic Data Integration, Harmonisation & Analytics
Day Three | Track 1: NGS Applications in the Clinic for Diagnostics Development
Day Three | Track 2: Multi-Omics in Cancer Discovery in Pharma
12th Annual Single Cell & Spatial Analysis UK Congress
This focused event brings together leading single cell analysis and spatial biology experts for 2-days of cutting-edge discussions, engaging roundtables, topical panel discussions, in-depth workshops, and high-level presentations covering the latest emerging tools, technologies, applications, and case studies. From new insights to the latest products and services, join us to cover topical bioinformatics updates & clinical development success stories.
- Hear from and meet with the key innovators in single cell & spatial multi-omics representing internationally renowned academic institutions, research institutes and hospitals as well as global pharmaceutical organisations and leading biotech companies.
- Discover collaborative solutions in novel single cell genomic, transcriptomic, proteomic, and metabolic technologies. Presentations will include new techniques for single cell imaging, one pot assays, and scalable single cell analysis pipelines for drug development
- Discuss the latest innovations pushing spatial technologies into the clinic. Take a deep dive into digital pathology, developing robust spatial markers, and AI-guided bioinformatics approaches
- Join sessions focused on key topics in single cell & spatial analysis: from benchmarking single cell technologies and single cell multi-omics through to dissecting population dynamics using single cell analyses and multiplexed spatial phenotyping
Agenda at a Glance
Day Two | Track 1: Moving Towards the Use of Spatial Technologies in the Clinic
Day Two | Track 2: Emerging Technologies & Tools In Single Cell Analysis
Day Three | Track 1: Applications of Spatial Technologies & Bioinformatics
Day Three | Track 2: Applications of Single Cell Omics in Humans
6th Annual Digital PCR Congress
An intensive two-day meeting that will delve into the latest advancements in, and applications of, digital PCR technologies from integration to the utilisation of liquid biopsies in the management, detection, and treatment of cancer. Bringing together like-minded experts and facilitating meaningful collaborations & partnerships in the field, this is the event to attend & join representatives from leading academic & research institutions and organisations to stay at the forefront of research.
- Hear from and meet with the key innovators in digital PCR & liquid biopsies. 2023 attendees included: Principal Investigator, World Health Organisation; Head of Molecular & Cellular Biology, MRC Harwell; and Associate Professor, Ghent University
- Explore the growing role of digital PRC in microbiology with sessions looking into the detection of infectious disease, anti-microbial resistance, and microbial quantification
- Discover collaborative solutions to Digital PCR advances, applications, integration & implementation. This focused event brings together key opinion leaders to discuss quality control of digital PCR assays and platforms, and optimisation of dPCR protocols to enhance diagnostic throughput
- Discuss the latest innovations in liquid biopsies. Presentations will delve into integrating liquid biopsies into clinical development, liquid biopsy workflows and advances in analysing cell-free DNA, CTCs, cell-free RNA, micro-RNAs
Agenda at a Glance
Day One | Track 5: Digital PCR in Oncology & Microbiology
Day Two | Track 5: Liquid Biopsies for Early Detection, Diagnosis & Disease Monitoring
CRISPR-based Medicine Congress
An intensive two-day meeting where groundbreaking advancements meet pioneering minds. From refined detection techniques to tailored delivery methods like viral vectors and nanoparticles, key opinion leaders will be exploring the tools driving CRISPR-based medicine forward. Discussions will also delve into the integration of CRISPR-based functional genomics in drug discovery, unveiling advanced screening approaches, novel therapeutic target identification, and the synergy with omics-based strategies. Bringing together like-minded experts representing academic institutions and pharmaceutical & biotech companies, the event serves as a platform to facilitate meaningful collaborations & partnerships in the field to stay abreast of current industry trends & challenges.
- Hear from and meet with the key innovators in CRISPR-based medicine. Join key opinion leaders from academic & research institutions as well as pharma & biotech companies to explore CRISPR-based medicine development – tools, delivery approaches and CRISPR-based functional genomics in drug discovery
- Explore improved detection techniques and choosing the right genome editing approach whilst also taking a deep dive into targeted delivery approaches & the development of gene therapies
- Discover collaborative solutions to CRISPR-based functional approaches in drug discovery. Pharma leaders will be covering advances in screening approaches, functional genomics in disease modelling and identification and characterisation of novel therapeutic targets
- Discuss the latest innovations in CRISPR-based medicine. Presentations will be covering computational tools and algorithms to design gRNAs for the CRISPR/Cas9 gene-editing systems, precision gene editing and CRISPR methods for genetic screens
Agenda at a Glance
Day One | Track 6 - CRISPR-based Medicine Development: Tools & Delivery
Day Two | Track 6 - CRISPR-based Functional Genomics in Drug Discovery





Networking & knowledge-sharing is at the heart of what we do. Alongside the innovative programme, attendees can engage in a variety of event features and make the most of participation in a range of experiences
-
Over 100 dynamic interactive discussions, roundtables, and presentations curated with insights from top industry leaders and experts
-
Global Technological Showcase
An international exhibition showcasing cutting-edge technologies and services from leading providers in the scientific field. -
Networking Emphasis
Unparalleled networking opportunities, including speed networking sessions, a vibrant drinks reception, and informal gatherings to foster meaningful connections -
Innovative Poster Displays
Inspiring poster displays and presentations unveiling the latest breakthroughs from emerging biotech, pharma, and academic institutions
Companies Represented Include











Inspiring Tomorrow's Breakthroughs: Unleashing Innovation & Collaboration in NextGen Omics Research, Development & Applications
Join engaging presentations featured in our innovation and collaboration tracks, inspiring growth and creating space for collaboration opportunities for all who attend. The core attendee mix includes early-stage and emerging biotech or (academic) spinoff companies developing their own platforms or technologies with fewer than 30 employees.
The Innovation & Collaboration track will be comprising of 10-minute presentations and focused panel discussion sessions, encouraging networking, facilitating collaborations & showcasing emerging innovations in nextgen omics research, development & applications.
Featuring Attendees from Biotech and/or (Academic) Spinoff Companies Interested In:
- Seeking collaboration partners with pharma and solution providers
- Discovering new investment opportunities with pharma
- Learning about the latest industry trends & challenges
- Keeping updated with new innovations and technology solutions
- Showcasing their company profile to a targeted and engaged audience
The NextGen Omics Young Scientist Awards include the best poster presentation award and are intended to honour an outstanding individual performance for a scientific work by a PhD student, PostDoc or early career scientist.
Please submit your poster presentation by no later than 25th September 2024 in the below categories:
- NGS & Clinical Diagnostics
- Single Cell & Spatial Analysis
- Precision Medicine
- CRISPR Applications & Editing
- DPCR & Liquid Biopsies
- Microbiome
- Student (PhD), PostDoc or early career scientist under 35 years of age. Proof must be provided via an active student ID card or a copy of status
- Applicants must be the first AND presenting author of the submitted paper and register for the meeting by 25th September 2024
- ONLY ONE YSA submission per person will be accepted. If authors submit multiple abstracts for consideration for the YSA, only one abstract will be taken into consideration for the YSA
Applicants must follow the procedure as follows:
- Register and submit an abstract using the registration link to the left on this page
- Once registered you will be provided with the online abstract submission form
- Prepare the poster or platform and present it (The poster presenter should be at the poster during all breaks.)
- The 3 winners with the highest scores will be announced at the drinks reception on the 23rd October 2024
- The winners’ 10-minute oral presentations will take place on the 24th October 2024.
Applicants must follow the procedure as follows:
- Register and submit an abstract by 25th September 2024 using the registration link to the left on this page
- Once registered you will be provided with the online abstract submission form
- Prepare the poster or platform and present it (The poster presenter should be at the poster during all breaks.)
- The 3 winners with the highest scores will be announced at the drinks reception on the 23rd October 2024
- The winners’ 10-minute oral presentations will take place on the 24th October 2024.
The platform or poster presentation will be evaluated by senior scientists (consisting of our speaking and steering group committee for the Omics series) on the basis of originality of the approach and quality of the work (e.g. appropriate methodology, interpretation of results, conclusiveness). All participants will receive the scoring and comments after the annual meeting via email.
The 3 winners receiving the highest scores will be announced at the drinks reception on the 23rd October 2024 and will be given a trophy as well as £1,000 contribution towards travel costs. Oxford Global will also provide winners with Omics PLUS Pass – 12 months access to Omics Series events and PLUS Pass online platform.
For further information contact help@oxfordglobal.com
Meet Our Expert Speakers
NextGen Omics 2024 again brings together a panel of prominent leaders and scientists, sharing new case studies, innovative, data and industry outlooks.
Below are some of our featured speakers:







All Speakers















.jpg?length=600&name=1dbb6196-e38a-4d74-bb68-d12f5babdb19-2-11764368speaker_photo-AIAS-Fellows-040219-SK04bw-web-Martin-Thomsen-(2).jpg)
















Meet Our Esteemed Sponsors
Diamond Sponsor
Platinum Sponsors

Gold Sponsors

Silver Sponsors

Bronze Sponsors
Network & Programme Sponsors
Interested in Sponsoring?
Oxford Global’s range of sponsorship opportunities are highly effective in enabling market immersion and visibility, with direct access to clients and potential customers.
Learn moreLatest Resources
Pianno: a Model for the Annotation of Spatial Transcriptomics Data

Harmonized Multi-Omics Data Analysis - How Does It Impact Pharmaceutical Research?

Roadblocks to the Sharing of Large Multi-Omics Datasets

Post-Event Proceedings – NextGen Omics US 2023

Interview with Gabriela Kramer Marek, Associate Professor, Reader in Molecular Imaging at The ICR

Diagnostic Breakthroughs: How Multi-Modal Signatures in Healthcare are Transforming Diagnosis and Treatment
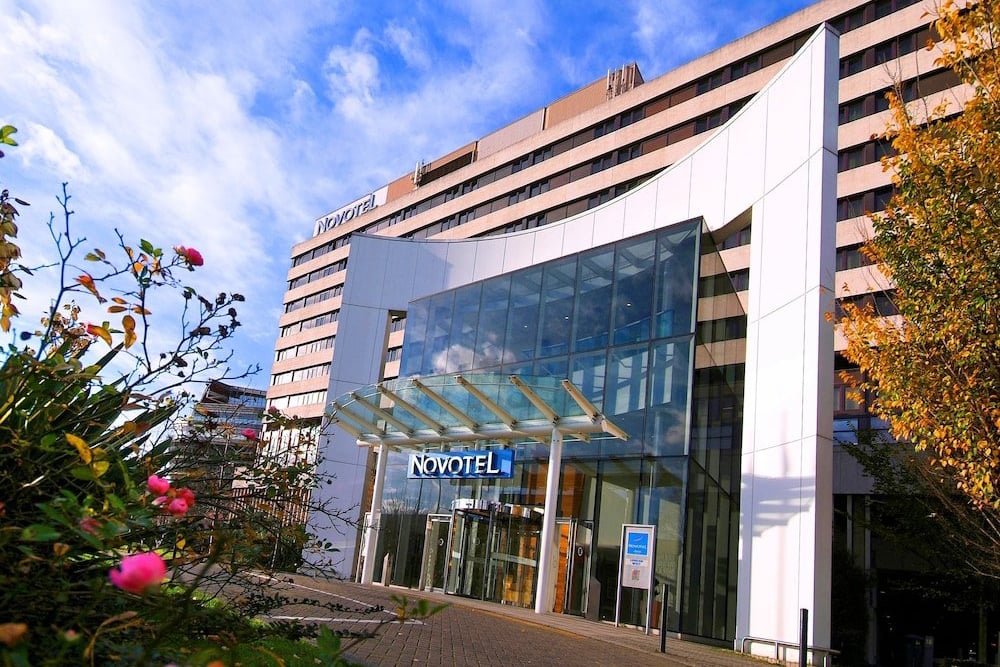
1 Shortlands,
Hammersmith International Ctre,
London, W6 8DR
Just minutes from three of London's main tube lines (Piccadilly, District and Hammersmith & City) and located in the heart of Hammersmith; Novotel London West is ideally located for trips to Westfield London, Harrods & Kensington High Street. Also conveniently located to Heathrow Airport with excellent road and rail links to the rest of the UK. This large and modern hotel offers on-site parking (chargeable), fitness suite and complimentary Wi-Fi throughout.
By Air
London Heathrow Airport - Novotel London West is accessible from Heathrow via the Underground on the Piccadilly Line - fares cost around £5 into Central London. A taxi from the airport will take approximately 20 minutes and will cost around £30 - £40.
London Gatwick Airport - The Gatwick Express runs every 15 mins - take it to Victoria station, and then get the District line to Hammersmith (about 15 mins). A taxi from the airport will take around 60 mins and cost between £65 - £80.
By Underground & Bus
Hammersmith Underground Station is adjacent to the hotel (3 minutes walk) with access to the Piccadilly, District and Hammersmith & City Lines. When exiting Hammersmith station, turn right and walk across the bus station. Cross over the roads using the island and keep on the right-hand side of Hammersmith Road. Continuing walking for 2 mins, and the hotel is accessible via stone steps.
For buses in central London, take route numbers 9 and 10. The main coach station (London Hammersmith) is 3 minutes walk away.
By Rail
The closest National Rail train station is Kensington Olympia (20 minutes walk).
By Car
Leave the A4 at the Hammersmith turning and proceed along Hammersmith Bridge Road to the large roundabout underneath the flyover. Take the fifth exit off the roundabout. Then turn left into Shortlands - the main hotel entrance and parking will be on your left-hand side.
Parking
Novotel London West offers over 240 on-site car parking spaces (charged per hour for residents parking). For further information including a map and full directions, please visit: www.novotel.com/gb/hotel-0737-novotel-london-west/index.shtml
We are closely monitoring the official guidance from health authorities, local governments, and the World Health Organization in order to support the health and well-being of our global community. The health and safety of our staff, customers and clients remains our number one priority.
As we continue to move forward with hosting our events in-person in 2024, we’ve added a series of Health & Safety guidelines and precautions in order to prepare for event safety. We carry out risk assessments for all our events to evaluate fundamental considerations and how to cover multiple risk scenarios.
Oxford Global has learned that third-party companies (recently EHotel Services, Business Travel Management/btravelmanagement and Exhibitors Hotel Reservations Services) are targeting conference attendees with a fraudulent hotel booking scheme.
Please note that none of these third-party companies are associated with Oxford Global in any way, nor have Oxford Global authorised them to use their names or trademarks on information they send out to attendees.
If you are contacted by a third-party company by phone or email using Oxford Global’s name or the name of NextGen Omics 2024 and offering accommodation services, we urge exhibitors and attendees to proceed with extreme caution before signing anything sent by these companies or entering into any conversation or replying to any emails sent from these third-party companies.
Register Your Interest
Submit your details and a member of our team will be in touch














.png?length=600&name=472937f4-077e-4fe7-bcb6-e7f6d0cc2621-2-5668375logo-Watchmaker_Logo_RGB_800px-(1).png)








